This document introduces several auxiliary functions of SigBridgeR.
Setting
Setting and Retriving Package Options
Currently, 3 package options are provided:
-
verbose: Whether to print the progress of the function. Default isTRUE. -
seed: The random seed used in the function. Default is123L. -
timeout: The maximum timeout time when downloading data. Default is180L.
These options can be modified using either the
setFuncOption or Options function (the former
is recommended).
setFuncOption(seed = 321L)
options(SigBridgeR.seed = 321L)setFuncOption automatically detects the prefix, so
parameters with a package name prefix are also compatible.
setFuncOption(SigBridgeR.seed = 321L)getFuncOption is used to retrieve the global parameters
of SigBridgeR.
getFuncOption("seed")
# 321
# * Auto-detect prefix
getFuncOption("SigBridgeR.seed")
# 321Setting Threads
setThreads() configures thread counts for OpenMP,
data.table, and TensorFlow backends. By default, it uses half of
available physical cores to balance performance and system
responsiveness.
Basic Usage
# Set explicit thread count
setThreads(8L)
# Configure specific backends only
setThreads(dt = 4L) # data.table only
setThreads(openmp = 4L) # OpenMP onlyTensorFlow Optimization
⚠️ Must configure BEFORE importing TensorFlow:
setThreads(
threads = 8L,
tf_config = list(
inter_op = 2L,
intra_op = 8L
)
)
tf <- reticulate::import("tensorflow") # Import AFTER configurationOptions
-
verbose = FALSE: suppress console output - Returns invisible list of old/new values for programmatic use
- Additional
data.table::setDTthreads()arguments accepted via...(e.g.,restore = TRUE)
💡 Tip: Default half-core allocation prevents oversubscription. Use full cores (
threads = availableCores()) only on dedicated compute nodes.
Seurat Utilities
Find the Optimal Normalization Method for Seurat
A simple scoring function for selecting a normalization method.
Evaluates 3 dimensions of preprocessing quality:
Variance stabilization: Decoupling of mean-variance relationship in normalized expression (lower correlation = better).
Biological signal retention: Preservation of known marker genes within highly variable genes (higher retention = better).
Dropout robustness: Removal of technical dropout bias from normalized values (lower correlation with dropout rate = better).
Exmaple Usage:
obj <- Seurat::CreateSeuratObject(counts)
sct <- Seurat::SCTransfrom(obj)
lognorm <- Seurat::NormalizeData(obj) %>%
Seurat::ScaleData() %>%
Seurat::FindVariableFeatures()
ChooseNormalization(
sct = sct,
lognorm = lognorm,
# * More can be added here
subset_size = integer(),
known_hvgs = list(),
n_hvgs = 2000L,
low_expressed_thresh = 0.2,
weight = c(
variance_stability = 0.4,
marker_signal = 0.35,
dropout_robustness = 0.25
)
)
# ℹ Using 1088 cells
# ℹ Comparing 2 methods: sct and lognorm
# ── Method Ranking (Composite Score)
# Top method: log_norm
# [1] log_norm: 0.825
# [2] sctransform: 0.175Supports simultaneously adding Seurat objects preprocessed with
multiple methods, ensuring they are equivalent to the standard workflow
of NormalizeData + ScaleData +
FindVariableFeatures.
Parameters
-
subset_size: Size of the single-cell data subset (i.e., number of cells) to be evaludated. -
known_hvgs: list or character. Known highly variable genes, provided as a reference baseline. -
n_hvgs: Maximum number of highly variable genes to consider. -
low_expressed_thresh: Percentage of genes excluded from consideration, ranked by expression level. -
weight: Scoring weight that sum to 1.
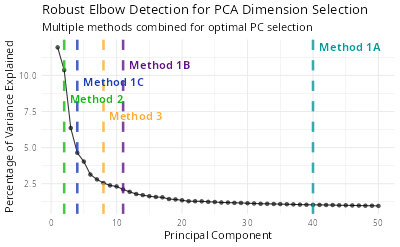
Finding the Optimal Number of Principle Components
Usually the number if principle components (PCs) is manually set to 10 or 20 according to the elbow plot. However, it is not always the case that the PCs with the highest variance are the most informative. Here is a function to help you find the optimal number of PCs.
ndims <- FindRobustElbow(
obj = seurat,
verbose = TRUE,
ndims = 50
)
# ── Method Results
# ℹ Method 1A (Cumulative Variance > 90%): 1:40
# ℹ Method 1B (Variance > Mean): 1:11
# ℹ Method 1C (Variance > 2*SD): 1:4
# ℹ Method 2 (Second Derivative): 1:2
# ℹ Method 3 (Distance-based): 1:8
# ✔ Final Recommended Dimensions: 1:35
# ── Summary
# ℹ Cumulative variance at 35 PCs: 84.8%
# ℹ Variance explained by PC35: 1.09%
print(ndims)
# 35This function will draw an elbow plot with each method and the
recommended number of PCs. You can also use the ndims
parameter to specify the maximum number of PCs to be tested. The default
value is 50.
knitr::include_graphics("vignettes/example_figures/elbow.png")Add miscellaneous information to the Seurat object
SigBridgeR uses AddMisc() to record what data features
or evidence the various screening algorithms are based on during
execution. As a substitute for the SeuratObject::Misc()
-
AddMisc(): Add miscellaneous information to the Seurat object. Support for adding multiple attributes to theSeuratObject@miscslot simultaneously.
# basic usage
seurat_obj <- AddMisc(seurat_obj, "QC_stats" = qc_df)
# Auto-incrementing example when `cover` set to FALSE
seurat_obj <- AddMisc(seurat_obj, markers = markers1)
seurat_obj <- AddMisc(seurat_obj, markers = markers2, cover=FALSE)
# Add multiple attributes to the `SeuratObject@misc` slot simultaneously
seurat_obj <- AddMisc(seurat_obj, markers1 = markers1, markers2 = markers2)
# Add multiple attributes (stored as a list element) to the `SeuratObject@misc`
seurat_obj <- AddMisc(seurat_obj, list(attr1 = "value1", attr2 = "value2"))Add Gene-level Metadata to the Seurat object
SigBridgeR uses AddMetaFeature() to update the
SeuratObject@assays$RNA@meta.data
Add a new column to the metadata
gene_type <- rep("test", nrow(seurat_obj))
seurat_obj <- AddMetaFeature(seurat_obj, "gene_type" = gene_type)Add a data.frame to the metadata, with the column names as the metadata names.
seurat_obj <- AddMetaFeature(
seurat_obj,
data.frame(gene_type = gene_type, gene_name = rownames(seurat_obj)
)Add to different assays
seurat_obj <- AddMetaFeature(seurat_obj, "gene_type" = gene_type, assay = "RNA")
seurat_obj <- AddMetaFeature(seurat_obj, "gene_type" = gene_type, assay = "ATAC")If duplicate column names are detected, they will be suffixed with an
underscore and a number (e.g., _1, _2) for
disambiguation.
Integration of Single Cell Data
Matrix Integration
When passing raw matrices, the function performs a “join” operation based on the union of all genes. Missing values are filled with NA.
# Create dummy matrices
mat1 <- matrix(rpois(50, 5), nrow = 10, dimnames = list(paste0("G", 1:10), paste0("C", 1:5)))
mat2 <- matrix(rpois(60, 6), nrow = 10, dimnames = list(paste0("G", 5:14), paste0("C", 1:6)))
# Integrate matrices with custom prefixes
integrated_mat <- SCIntegrate(BatchA = mat1, BatchB = mat2)
# default prefixed
integrated_mat2 <- SCIntegrate(mat1, mat2)
integrated_mat[1:6, 1:4]
# BatchA_C1 BatchA_C2 BatchA_C3 BatchA_C4
# G1 3 2 4 5
# G10 2 4 3 4
# G11 NA NA NA NA
# G12 NA NA NA NA
# G13 NA NA NA NA
# G14 NA NA NA NA
integrated_mat2[1:6, 1:4]
# mat1_C1 mat1_C2 mat1_C3 mat1_C4
# G1 3 2 4 5
# G10 2 4 3 4
# G11 NA NA NA NA
# G12 NA NA NA NA
# G13 NA NA NA NA
# G14 NA NA NA NAKey Features:
Duplicate Resolution: Automatically calls
AggregateDupsto handle redundant gene symbols.Auto-Naming: Uses argument names (like BatchA) as cell ID prefixes.
Seurat Integration
For Seurat objects, SCIntegrate automates the standard
workflow via the pipeline parameter.
The Pipeline String: Each letter in the pipeline argument triggers a specific Seurat command:
| Code | Function | Description |
|---|---|---|
| o | CreateSeuratObject |
(do not use it in this function) |
| n | NormalizeData |
Standard normalization. |
| s | ScaleData |
Scales data for PCA. |
| v | FindVariableFeatures |
Selects highly variable genes. |
| p | RunPCA |
Principal Component Analysis. |
| e | FindNeighbors |
Computes SNN graph. |
| c | FindClusters |
Louvain algorithm clustering. |
| t | RunTSNE |
t-SNE reduction. |
| u | RunUMAP |
UMAP reduction. |
| r | SCTransform |
SCT workflow. Replaces n, s, v. |
If more are needed, see A Guide for Custom Extensions
Example Usage
integrated <- SCIntegrate(
obj1, obj2, # -> merge.Seurat
pipeline = "nsvpi",
method = Seurat::RPCAIntegration, # Change integration method
dims = 1:30, # Passed to RunPCA and IntegrateLayers
k.weight = 50 # Passed to IntegrateLayers
)An example using mock data
mat1 <- matrix(
rpois(1000, 5),
nrow = 20,
dimnames = list(paste0("G", 1:20), paste0("C", 1:50))
)
mat2 <- matrix(
rpois(1200, 6),
nrow = 20,
dimnames = list(paste0("G", 11:30), paste0("C", 1:60))
)
seu1 <- Seurat::CreateSeuratObject(mat1)
seu2 <- Seurat::CreateSeuratObject(mat2)
integrated_seu <- SCIntegrate(
Seurat1 = seu1, # Add prefixes
Seurat2 = seu2,
method = Seurat::CCAIntegration,
pipeline = "nsvpi",
dims = 1:10,
k.weight = 40
)
integrated_seu
# An object of class Seurat
# 30 features across 110 samples within 1 assay
# Active assay: RNA (30 features, 10 variable features)
# 5 layers present: counts.1, counts.2, data.1, data.2, scale.data
# 2 dimensional reductions calculated: pca, integrated.drCell Type Annotation
As of version 3.3.0, three built-in cell annotation algorithms are included and can be applied directly to Seurat objects. Using them requires installing additional dependencies. See the README.
Basic usage:
seurat <- SCAnnotate(
sc = seurat,
method = c("CellTypist", "SingleR", "mLLMCelltype")
# * more arguments passed to each method
)SCAnnotate will automatically set up the required
dependencies for you (making it even more plug-and-play). If the
method argument is not specified, it will automatically
select the appropriate annotation method based on the provided
arguments.
To control it manually, use following functions:
CellTypist Annotation
This is the direct R wrapper of celltypist.annotate. Unlike
SCAnnotate, it does not prepare an environment for you.
seurat <- CellTypistAnnotate(
sc,
model = NULL,
download = TRUE,
conda = NULL,
python = NULL,
verbose = getFuncOption("verbose"), # TRUE
celltypist_tools = system.file(
"python/73-CellTypistAnnotate.py",
package = "SigBridgeR"
),
# * more arguments pass to celltypist.annotate (python)
) Parameters:
-
sc: A Seurat object. -
model: Charater/str. The model to use. IfNULL, the default model will be used. -
download: Logical/bool. Whether to download the model. Defaults toTRUE-download all models. if model already exists, skip download. -
force_update: Logical. Whether to force update the model file. -
conda: Character/str. conda environment name. IfNULL, try to usepython. -
python: Character/str. The path to the python executable. IfNULL, try to useconda. -
verbose: Logical/bool. Whether to print verbose messages. Defaults toTRUE. -
celltypist_tools: The path to the celltypist_tools.py file. IfNULL, the default path will be used.
Returns:
A Seurat object with the cell type annotation added as metadata columns.
Other parameters supported by celltypist.annotate will
be automatically passed through and can be directly supplied to
CellTypistAnnotate.
helpful link: https://github.com/Teichlab/celltypist
mLLMCelltype Annotation
seurat <- mLLMCellTypeAnnotate(
sc,
seurat_obj_markers = NULL,
# * Context for large language models
tissue_name = "Human Cancer",
models = c(
"gpt-5",
"claude-sonnet-4-5-20250929",
"gemini-3-pro",
"qwen-max-2025-01-25"
),
api_keys = list(
anthropic = "your-anthropic-key",
openai = "your-openai-key",
gemini = "your-google-key",
qwen = "your-qwen-key"
),
# Other arguments passed to `mLLMCelltype::annotate_cell_types` or ` mLLMCelltype::interactive_consensus_annotation`
)Parameters
sc: A Seurat object with pre-computed clusters (stored inIdents(sc)orsc$seurat_clusters).seurat_obj_markers: Optional pre-computed marker gene table or list (output ofSeurat::FindAllMarkers()). IfNULL(default), markers are computed automatically using parameters passed via....tissue_name: Character. Biological context for annotation (e.g., tissue type, disease state). Helps LLMs interpret marker genes appropriately. Default:"Human Tumor".-
models: Character vector of LLM model identifiers. Supported formats:- OpenAI:
"gpt-4o","gpt-4o-mini", etc. - Anthropic:
"claude-3-5-sonnet-20240620", etc. - Google:
"gemini-1.5-pro", etc. - Alibaba:
"qwen-max","qwen-plus", etc.
Default:
c("gpt-4o", "claude-3-5-sonnet-20240620", "gemini-1.5-pro", "qwen-max"). For single-model mode, only the first model is used. - OpenAI:
-
api_keys: Named list of API keys with provider names as keys:-
openai: OpenAI API key -
anthropic: Anthropic API key -
gemini: Google Cloud API key (with Gemini enabled) -
qwen: Alibaba DashScope API key
Example:
list(openai = "sk-...", anthropic = "sk-ant-...").Note: Default placeholder keys (
"your-xxx-key") will fail—users must supply valid keys. -
Returns:
A Seurat object with the cell type annotation added as metadata columns.
helpful link: https://github.com/cafferychen777/mLLMCelltype
SingleR Annotation
seurat <- SingleRAnnotate(
sc,
verbose = getFuncOption("verbose"), # TRUE
# * pass to `celldex::HumanPrimaryCellAtlasData`
ensembl = FALSE,
cell.ont = c("all", "nonna", "none"),
legacy = FALSE,
# * pass to `SingleR::SingleR`
ref = c("HPCA", "custom"),
labels = NULL,
# Other arguments passed to `SingleR::SingleR`
)Parameters
-
sc: A Seurat object. -
verbose: Logical. Whether to print progress messages. Defaults toTRUE. -
ensembl: Logical. Whether to convert row names to Ensembl IDs. Genes without a valid mapping are discarded. -
cell.ont: Character. specifying Cell Ontology term handling. Use"nonna"to discard samples without valid terms,"all"to keep all samples (with possibleNA), or"none"to skip. -
legacy: Logical. Whether to use legacy ExperimentHub backend. Default uses gypsum. -
ref: Reference data for annotation. Either"HPCA"(Human Primary Cell Atlas fromcelldex) or aSingleCellExperimentobject with pre-labeled cells. For multiple references, a list of matrices orSummarizedExperimentobjects. -
labels: Character vector of cell type labels forrefcolumns. Required for custom references; defaults toref$label.mainfor"HPCA". Ifrefis a list,labelsmust be a corresponding list.
Returns:
A Seurat object with the cell type annotation added as metadata columns.
helpful link: https://github.com/SingleR-inc/SingleR
Setting up Python Environment
Some screening methods (e.g. Section 3.5 DEGAS) are built using Python and require an execution environment. Here is a function to help you set up a Python environment. Both Windows and Unix-like systems are supported.
# * This is an example of setting up a Python environment using conda
SetupPyEnv(
env_type = "conda",
env_name = "test-condaenv",
method = c("reticulate", "system","environment"), # choose one of them, default is "reticulate"
env_file = NULL # path to environment.yml file, used when method = "environment"
python_version = NULL,
packages = c(
"pandas" = "1.3",
"numpy" = "any"
),
recreate = FALSE, # whether to remove the existing environment and recreate it
use_conda_forge = TRUE,
verbose = TRUE
)
reticualte::use_condaenv("test-condaenv")
# * Or use virtualenv via reticulate
SetupPyEnv(
env_type = "venv",
env_name = "test-venv",
python_version = "3.9.15",
packages = c("tensorflow" = "2.4.1", "protobuf" = "3.20.3"),
python_path = NULL,
recreate = FALSE,
verbose = TRUE
)
reticulate::use_virtualenv("test-venv")You can use ListPyEnv() to list all the Python
environments you have set up. Both conda and virtual environments are
supported.
# * Unix-like systems
ListPyEnv()
# name python type
# 1 base /home/user/miniconda3/bin/python conda
# 2 test-condaenv /home/user/miniconda3/envs/test-condaenv/bin/python conda
# 3 test-venv /home/user/miniconda3/envs/test-venv/bin/python venvShow the conda environments only:
# * Unix-like systems
ListPyEnv(env_type = "conda")
# name python type
# 1 base /home/user/miniconda3/bin/python conda
# 2 test-condaenv /home/user/miniconda3/envs/test-condaenv/bin/python condaIf the virtual environment isn’t installed in the default location,
you can specify the location of the virtual environment with the
venv_locations parameter.
ListPyEnv(env_type = "venv", venv_locations ="~/here_is_a_dir/.virtualenvs")Other Functions
Load reference data
Parameters
-
data_type: The type of data to load. Can be either “continuous”, “survival” or “binary”, case-insensitive. -
path: The path to the data directory. -
cache: Whether to cache the data. Defaults toTRUE. -
timeout: The maximum timeout time when downloading data.
When loading the example data, the single-cell RNA expression matrix, the bulk RNA expression matrix, and the clinical phenotype data are loaded all at once. These data are combined into a list and returned.
mydata <- LoadRefData(
data_type = c("survival"),
path = tempdir(),
cache = TRUE,
timeout = 60
)
# * mat_exam (matrix_example)
mydata[[1]][1:6,1:6]
# SMC01.T_AAACCTGCATACGCCG SMC01.T_AAACCTGGTCGCATAT SMC01.T_AAACCTGTCCCTTGCA SMC01.T_AAACGGGAGGGAAACA SMC01.T_AAACGGGGTATAGGTA SMC01.T_AAAGATGAGGCCGAAT
# A1BG 0 0 0 0 0 0
# A1BG.AS1 0 0 0 0 0 0
# A1CF 0 2 0 0 3 0
# A2M 0 0 0 0 0 0
# A2M.AS1 0 0 0 0 0 0
# A2ML1 0 0 0 0 0 0
# * bulk_survival
mydata[[2]][1:6,1:6]
# TCGA-69-7978 TCGA-62-8399 TCGA-78-7539 TCGA-73-4658 TCGA-44-6775 TCGA-44-2655
# HIF3A 4.2598 11.6239 9.1362 5.0288 4.0573 5.5335
# RTN4RL2 8.2023 5.5819 3.5365 7.4156 7.7107 5.3257
# HMGCLL1 2.7476 5.8513 3.8334 3.6447 2.9188 4.8820
# LRRTM1 0.0000 0.4628 4.7506 6.8005 7.7819 2.2882
# GRIN1 6.6074 5.4257 4.9563 7.3510 3.5361 3.3311
# LRRTM3 1.7458 2.0092 0.0000 1.4468 0.0000 0.0000
# * pheno_survival
mydata[[3]] |> head()
# time status
# TCGA-69-7978 4.40 0
# TCGA-62-8399 88.57 0
# TCGA-78-7539 25.99 0
# TCGA-73-4658 52.56 1
# TCGA-44-6775 23.16 0
# TCGA-44-2655 43.50 0We recommend using the zeallot package’s
%<-% function to assign values and rename them
simultaneously.
Aggregating Duplicate Rows or Columns
These functions collapse duplicated row names (e.g., gene symbols) or column names (e.g., sample IDs) in expression matrices and count tables using configurable aggregation methods.
Core Functions
| Function | Purpose |
|---|---|
AggregateDupRows() |
Merge rows with identical names (e.g., duplicate genes) |
AggregateDupCols() |
Merge columns with identical names (e.g., duplicate samples) |
AggregateDups() |
Convenience wrapper: merge rows & columns |
Basic Usage
# Example matrix with duplicate genes (rows) and samples (columns)
mat <- matrix(1:16, nrow = 4,
dimnames = list(c("TP53", "TP53", "BRCA1", "ACTB"),
c("S1", "S1", "S2", "S3")))
mat
# S1 S1 S2 S3
# TP53 1 5 9 13
# TP53 2 6 10 14
# BRCA1 3 7 11 15
# ACTB 4 8 12 16
# Collapse duplicate genes using sum
AggregateDupRows(mat, method = "sum")
# S1 S1.1 S2 S3
# TP53 2 6 10 14
# BRCA1 3 7 11 15
# ACTB 4 8 12 16
# Collapse duplicate samples using max
AggregateDupCols(mat, method = "max")
# S1 S2 S3
# TP53 5 9 13
# TP53 6 10 14
# BRCA1 7 11 15
# ACTB 8 12 16
# Full deduplication in one step
AggregateDups(mat, method = "sum")
# S1 S2 S3
# TP53 14 19 27
# BRCA1 10 11 15
# ACTB 12 12 16